Research
Microbial ecosystems in the high-diversity regime
My research lies at the intersection of statistical physics, classical ecology, experimental microbiology and bioinformatics. My primary focus is in investigating microbial ecosystems in the high-diversity regime. The traditional approach of building biological understanding ‘from the bottom up,’ seeking to explain the behavior of a system from the interactions of individual components. For natural microbial ecosystems that harbor hundreds of interacting ‘species’ such an approach is challenging, and laboratory investigations often focus on simplified systems. My group builds on the ideas of statistical physics to adopt the opposite, “top-down” approach, studying complex ecosystems directly in a naturalistic context, and identifying which mesoscopic properties, and under what conditions, are predictable. Our goals are (1) to identify the principles governing the complexity of the simplest ‘good-enough’ model for a given functional property of an ecosystem; (2) to develop theoretical and computational tools for learning task-dependent coarse-graining of microbial ecosystems from data; and (3) to harness this understanding for top-down design of functional communities.
Microbial systems present a fascinating theoretical challenge: The traditional ecological units such as ‘species’ are ill-defined. Two bacteria we call ‘E. coli’ can differ over half of their gene content, and we know that both community dynamics and function are shaped by details down to the level of individual strains. Thus, any variables we might realistically use in our equations are always collapsing relevant functional diversity. This seems like a monumental problem for modeling. And yet, methane production in a wastewater bioreactor can be predicted with a model coarse-graining its 1500+ species into just 4 functional classes. In other words, the species-level description is simultaneously too microscopic and not microscopic enough! When is it ok to ignore diversity? Which ecosystem properties are "coarse-grainable"? Understanding and leveraging coarse-grainability could transform our ability to predict and control microbial systems, and we are very excited to be investigating this both in our theory work and in partnership with experimental groups (Seppe Kuehn, UChicago; Otto Cordero, MIT).
Data-driven collaborations
The research in our group also has a second, independent, thrust: working with highly quantitative biological data to extract more out of it. Biological data is high-dimensional and complex, and sometimes messy—but many modern devices used to collect it (e.g. sequencers, microscopes) are incredibly accurate machines. This accuracy is often underexploited, with the reasoning that biological variability is too large to justify going beyond semi-quantitative methods. However, this perceived biological variability is often a consequence of biases or inaccuracies of the analysis itself. Forgoing canned statistical tools for visualizations and analyses custom-built for a given experiment, tailored to the strengths and limitations of a particular experimental protocol allows us to uncover biases and systematic errors, develop methods to correct them, and reveal unexpected signal modalities. My past work included single-molecule imaging and high-resolution analysis of genomic and sequencing data (which has some cool applications). Current collaborations in my group focus on correcting amplification bias in barcoding experiments (with D. Petrov, Stanford); high-resolution spectral imaging of cell organelles (with S. Mukherji, WashU) and high-resolution structure-to-function mapping of soil microbial ecosystems (with S. Kuehn, UChicago).
Big picture: the dreams
Video: Mikhail Tikhonov - “Microbial ecology as a new frontier for theoretical physics” (Stanford Physics/Applied Physics colloquium)
Beyond discreteness: "species" in the microbial world
Most of classical ecology relies on the assumption that organisms can be naturally partitioned into discrete types. But how well does this intuition actually apply to microbes? Asexual reproduction, evolution, horizontal gene transfer: all these mechanisms render the "partitioning assumption" questionable. It is clear that "species" is a useful concept. However, right now, this is not the best language we have, it is the only one we have – and many fascinating examples, like the stories of Methanobacillus omelianskii or Chlorochromatium aggregatum, make one wonder just how good this discrete language really is.
Unfortunately, no alternative currently exists. Our experimental studies report the "number of species" in the gut. Our models investigate dynamics of "species abundance", as shaped by "species interactions". Data analysis packages cluster sequences into "taxonomic units". Thus the partitioning assumption shapes our questions, our models, and even our approach to data. How else could we describe microbial ecosystems, if not in these terms?
The implications of high dimensionality
Our understanding of ecological and evolutionary mechanisms derives predominantly from analysis of low-dimensional models (with few species). However, natural communities are famously highly diverse, with hundreds of phenotypes coexisting and interacting. In physics, we know that when the number of degrees of freedom becomes large, the behavior of the system may change completely. What is qualitatively different about the high-diversity regime in ecology?
There exists a long tradition of "large-N ecology", starting with the classic work of Robert May. However, the focus has traditionally been on composition, e.g. the number of coexisting species. What are the specifically functional implications of high diversity? And how should this modify the familiar low-dimensional intuition we have of ecological and evolutionary processes?
Blurring the species boundaries
If research groups had mascots, we'd be proudly represented by the duck-rabbit!
Collaborators, past and current:
Joy Bergelson (NYU)
William Bialek (Princeton)
Michael Brenner (Harvard)
Otto Cordero (MIT)
Daniel Fisher (Stanford)
Hernan Garcia (Berkeley)
Thomas Gregor (Princeton)
Shamit Kachru (Stanford)
Seppe Kuehn (UChicago)
Pankaj Mehta (BU)
Remi Monasson (ENS Paris)
Sampriti Mukherjee (U Chicago)
Shankar Mukherji (WashU)
Dmitri Petrov (Stanford)
Alvaro Sanchez (Yale)
Justin Sonnenburg (Stanford)
Ned Wingreen (Princeton)
Statistical physics of resource competition
One classic way to link composition and function is provided by MacArthur's resource competition model from the late 60's. It was extensively studied for N = 1 and N = 2 resources, in particular by Tilman, who developed a highly influential geometric intuition.
In natural communities, however, the number of relevant metabolites is in the dozens. Remarkably, the high-diversity regime of the MacArthur model can be solved analytically, using methods of statistical physics of disordered systems. This exactly solvable model provides a rich platform to investigate the implications of high dimensionality for both ecological and evolutionary dynamics.
Moran J & Tikhonov M (2022). Defining coarse-grainability in a model of structured microbial ecosystems. PRX (in press)
Bergelson J et al. (2021) Functional biology in its natural context: A search for emergent simplicity. eLife
Goldford J, Lu N et al. (2018) Emergent simplicity in microbial community assembly. Science
Tikhonov M, Monasson R (2018) Innovation rather than improvement: a solvable high-dimensional model highlights the limitations of scalar fitness. J Stat Phys
Tikhonov M, Monasson R (2017) Collective phase in resource competition in a highly diverse ecosystem. PRL
Community-level competition
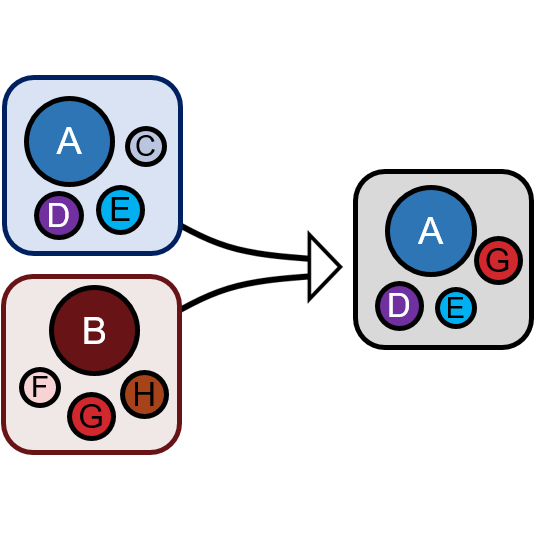
Natural microbial ecosystems do not just exchange members: they frequently come into contact as entire communities. For instance, think of a leaf falling on the ground, or a dog licking your hand. These events have been dubbed "community coalescence". A frequent, yet largely unstudied natural occurrence, they are expected to be an important factor shaping community structure, and are also of significant medical interest (e.g. fecal matter transplants are one of the few effective therapies against C. difficile infections). Interpreted as community-level competition, community coalescence is a very intriguing phenomenon also from a theoretical standpoint.
In collaboration with Alvaro Sanchez (Yale) and Pankaj Mehta (Boston U), we are investigating community coalescence both theoretically and experimentally.
Díaz-Colunga J, Lu N, Sanchez-Gorostiaga A et al (2022) Top-down and bottom-up cohesiveness in microbial community coalescence. PNAS
Tikhonov M (2016) Community-level cohesion without cooperation. eLife
Other projects
Other recent and ongoing projects cover a diverse set of topics, including:
regulatory circuits that can learn environment fluctuations (eLife, 2021);
tradeoffs of evolution (PRE, 2021) and evolution of tradeoffs (PNAS, 2020);
metabolically driven spatial assembly of multi-species consortia (PNAS, 2018);
"weakly structured" ecologies where the number of species is ill-defined (PRE, 2017).